geom_qq() and stat_qq() produce quantile-quantile plots. geom_qq_line() and
stat_qq_line() compute the slope and intercept of the line connecting the
points at specified quartiles of the theoretical and sample distributions.
Usage
geom_qq_line(
mapping = NULL,
data = NULL,
geom = "path",
position = "identity",
...,
distribution = stats::qnorm,
dparams = list(),
line.p = c(0.25, 0.75),
fullrange = FALSE,
na.rm = FALSE,
show.legend = NA,
inherit.aes = TRUE
)
stat_qq_line(
mapping = NULL,
data = NULL,
geom = "path",
position = "identity",
...,
distribution = stats::qnorm,
dparams = list(),
line.p = c(0.25, 0.75),
fullrange = FALSE,
na.rm = FALSE,
show.legend = NA,
inherit.aes = TRUE
)
geom_qq(
mapping = NULL,
data = NULL,
geom = "point",
position = "identity",
...,
distribution = stats::qnorm,
dparams = list(),
na.rm = FALSE,
show.legend = NA,
inherit.aes = TRUE
)
stat_qq(
mapping = NULL,
data = NULL,
geom = "point",
position = "identity",
...,
distribution = stats::qnorm,
dparams = list(),
na.rm = FALSE,
show.legend = NA,
inherit.aes = TRUE
)Arguments
- mapping
Set of aesthetic mappings created by
aes(). If specified andinherit.aes = TRUE(the default), it is combined with the default mapping at the top level of the plot. You must supplymappingif there is no plot mapping.- data
The data to be displayed in this layer. There are three options:
If
NULL, the default, the data is inherited from the plot data as specified in the call toggplot().A
data.frame, or other object, will override the plot data. All objects will be fortified to produce a data frame. Seefortify()for which variables will be created.A
functionwill be called with a single argument, the plot data. The return value must be adata.frame, and will be used as the layer data. Afunctioncan be created from aformula(e.g.~ head(.x, 10)).- geom
The geometric object to use to display the data, either as a
ggprotoGeomsubclass or as a string naming the geom stripped of thegeom_prefix (e.g."point"rather than"geom_point")- position
Position adjustment, either as a string naming the adjustment (e.g.
"jitter"to useposition_jitter), or the result of a call to a position adjustment function. Use the latter if you need to change the settings of the adjustment.- ...
Other arguments passed on to
layer(). These are often aesthetics, used to set an aesthetic to a fixed value, likecolour = "red"orsize = 3. They may also be parameters to the paired geom/stat.- distribution
Distribution function to use, if x not specified
- dparams
Additional parameters passed on to
distributionfunction.- line.p
Vector of quantiles to use when fitting the Q-Q line, defaults defaults to
c(.25, .75).- fullrange
Should the q-q line span the full range of the plot, or just the data
- na.rm
If
FALSE, the default, missing values are removed with a warning. IfTRUE, missing values are silently removed.- show.legend
logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes. It can also be a named logical vector to finely select the aesthetics to display.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.borders().
Aesthetics
stat_qq() understands the following aesthetics (required aesthetics are in bold):
Learn more about setting these aesthetics in vignette("ggplot2-specs").
stat_qq_line() understands the following aesthetics (required aesthetics are in bold):
Learn more about setting these aesthetics in vignette("ggplot2-specs").
Computed variables
These are calculated by the 'stat' part of layers and can be accessed with delayed evaluation.
Variables computed by stat_qq():
after_stat(sample)
Sample quantiles.after_stat(theoretical)
Theoretical quantiles.
Variables computed by stat_qq_line():
after_stat(x)
x-coordinates of the endpoints of the line segment connecting the points at the chosen quantiles of the theoretical and the sample distributions.after_stat(y)
y-coordinates of the endpoints.
Examples
# \donttest{
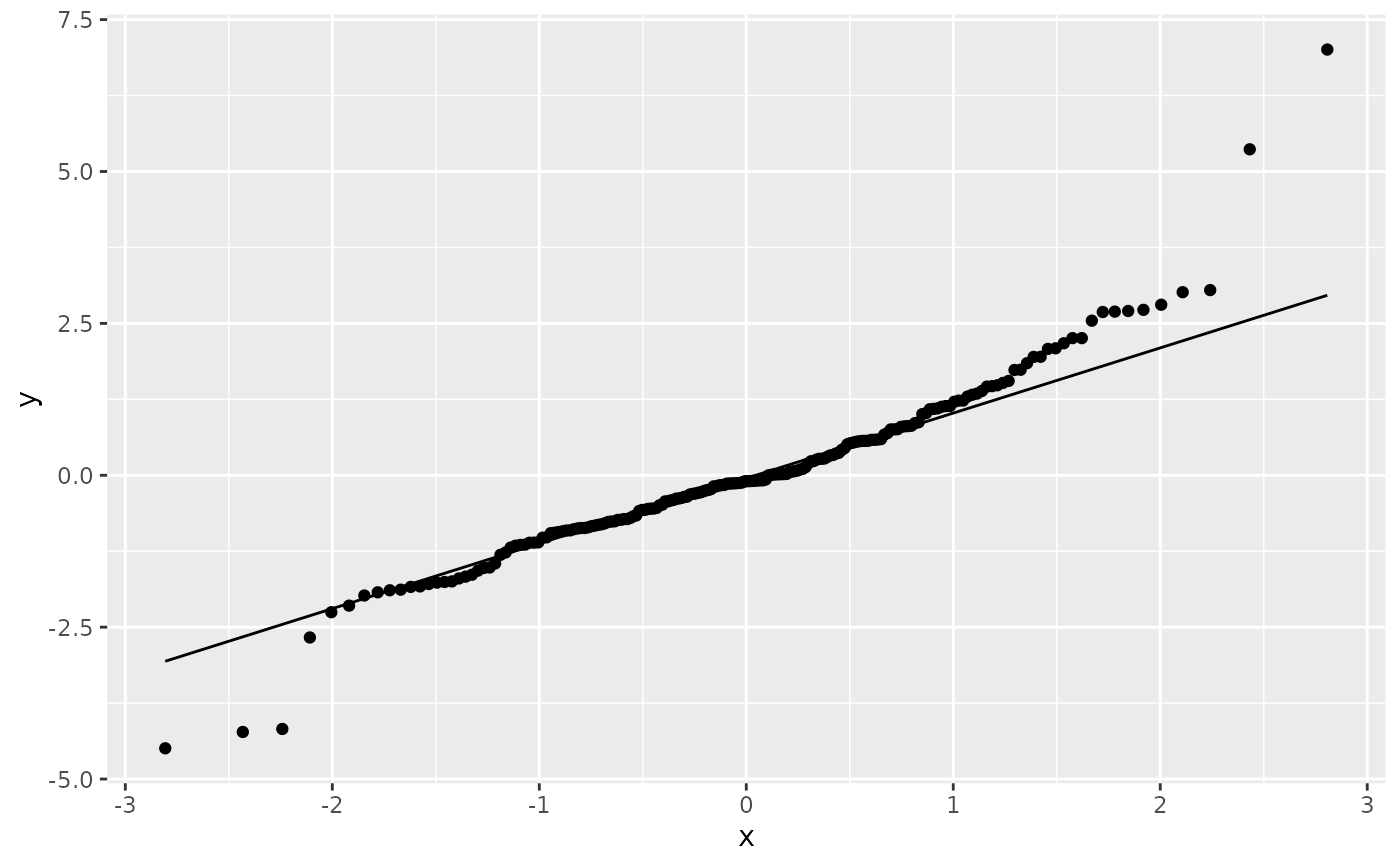
df <- data.frame(y = rt(200, df = 5))
p <- ggplot(df, aes(sample = y))
p + stat_qq() + stat_qq_line()
 # Use fitdistr from MASS to estimate distribution params
params <- as.list(MASS::fitdistr(df$y, "t")$estimate)
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
ggplot(df, aes(sample = y)) +
stat_qq(distribution = qt, dparams = params["df"]) +
stat_qq_line(distribution = qt, dparams = params["df"])
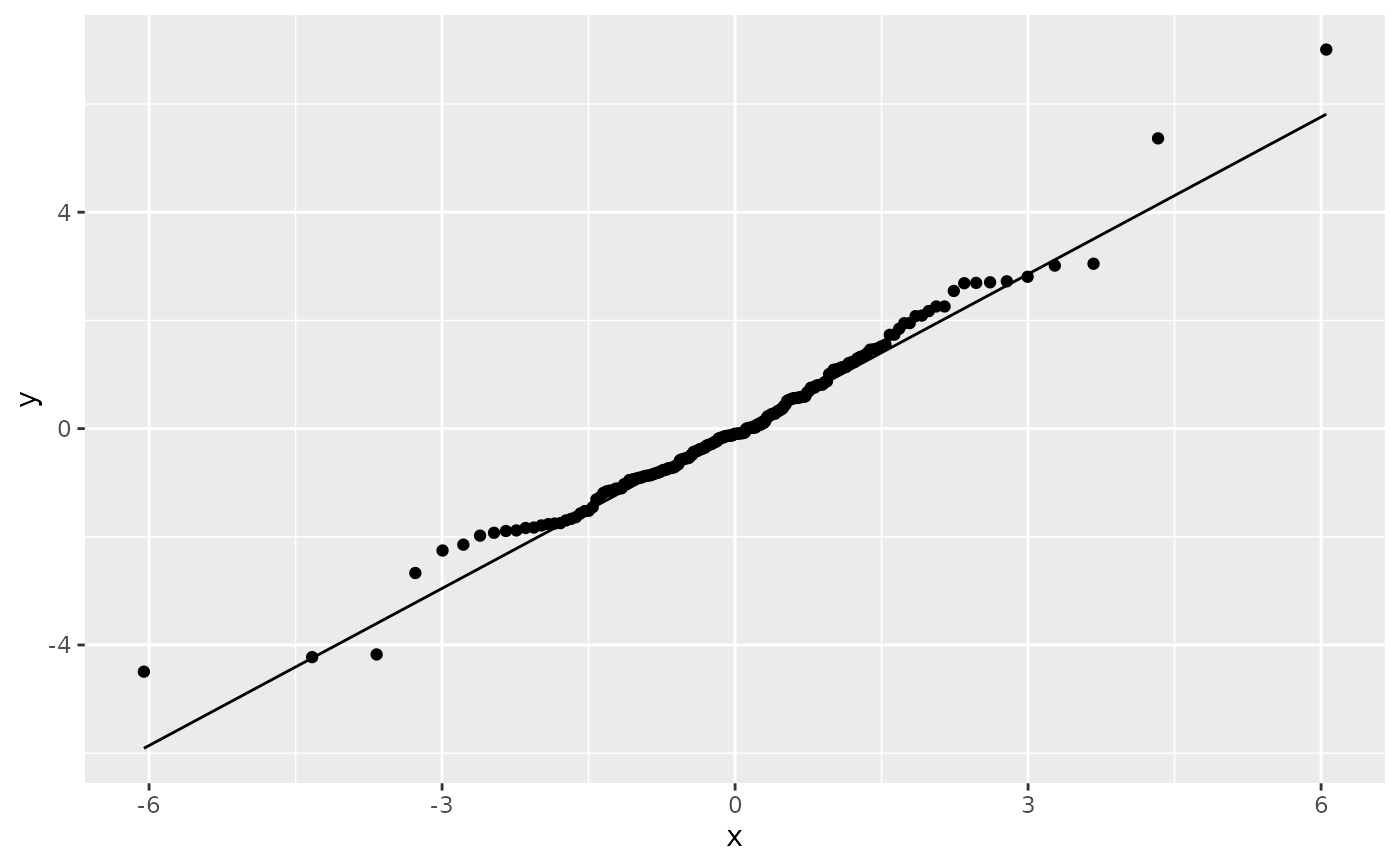
# Use fitdistr from MASS to estimate distribution params
params <- as.list(MASS::fitdistr(df$y, "t")$estimate)
#> Warning: NaNs produced
#> Warning: NaNs produced
#> Warning: NaNs produced
ggplot(df, aes(sample = y)) +
stat_qq(distribution = qt, dparams = params["df"]) +
stat_qq_line(distribution = qt, dparams = params["df"])
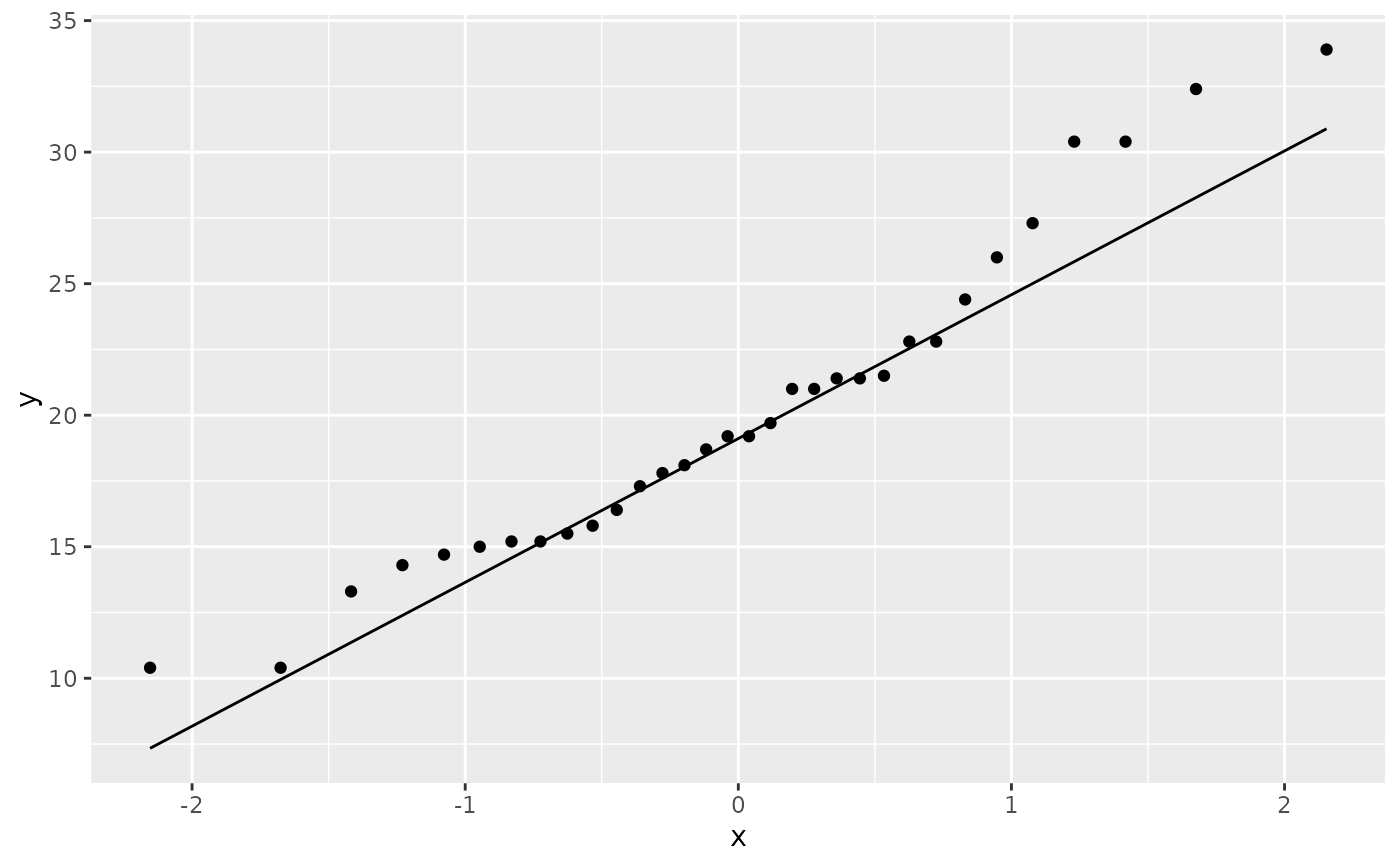
 # Using to explore the distribution of a variable
ggplot(mtcars, aes(sample = mpg)) +
stat_qq() +
stat_qq_line()
# Using to explore the distribution of a variable
ggplot(mtcars, aes(sample = mpg)) +
stat_qq() +
stat_qq_line()
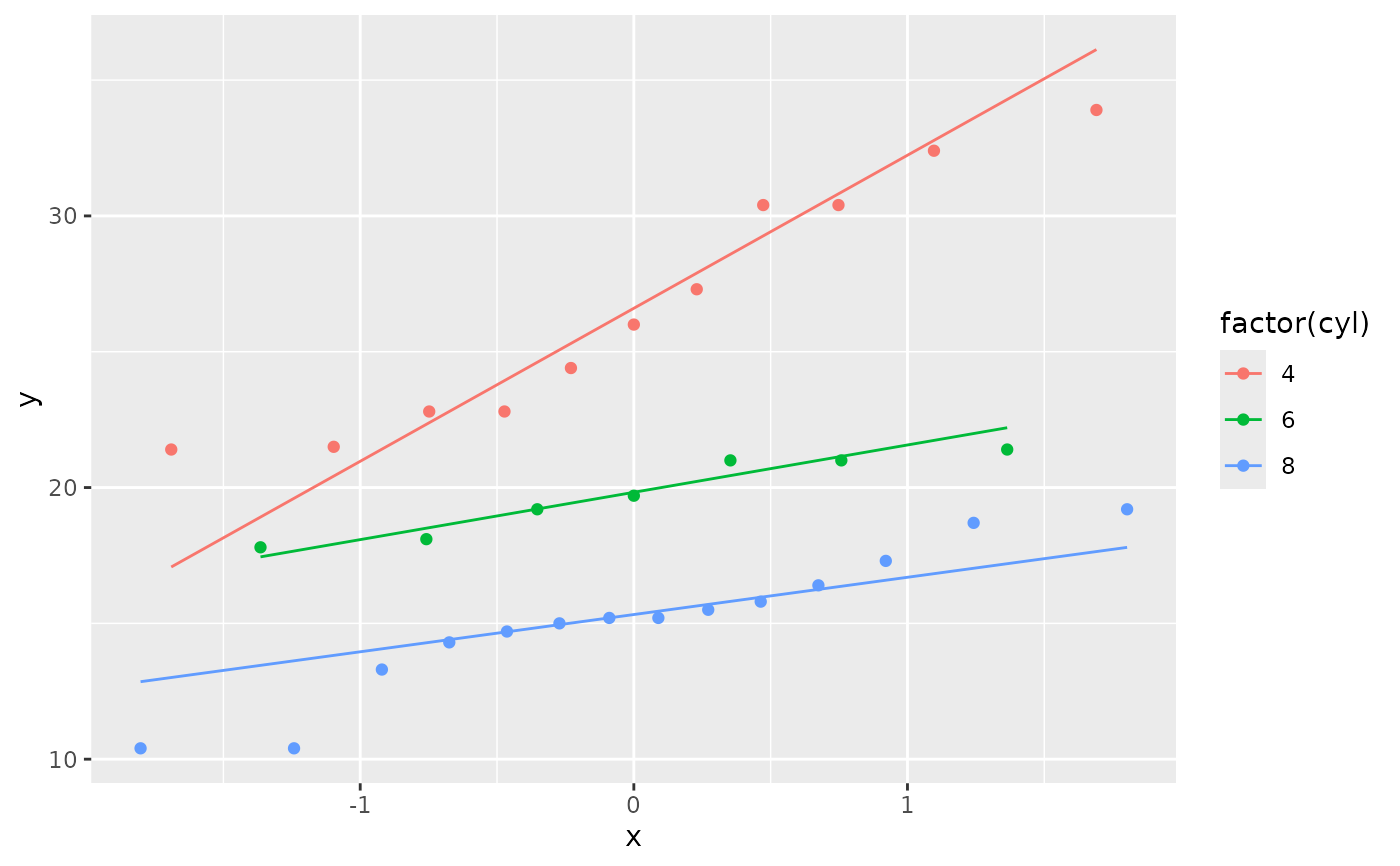
 ggplot(mtcars, aes(sample = mpg, colour = factor(cyl))) +
stat_qq() +
stat_qq_line()
ggplot(mtcars, aes(sample = mpg, colour = factor(cyl))) +
stat_qq() +
stat_qq_line()
 # }
# }
